In this post, we will be doing some reverse code engineering. This time not the same type of code we’re all familiar with, in today’s post we will be doing some Biological code Reverse Engineering, we will be reversing our first Genome, the Escherichia MS2 Bacteriophage.
The reason I chose E. MS2 specifically is that its Genome is one of the shortest sequences discovered until now. Also, I find it fascinating to study a Bacteriophage; a Virus that attacks Bacteria, How amazing that is!
The E. MS2 is a Virus that attacks E. Coli Bacteria which is one of the Bacterias that sits in an organisms intestine. But what happens when an innocent E. Coli Bacteria is infected with the E. MS2 Virus? That’s what we are supposed to answer in this post by taking apart the viral DNA and reversing its synthesized Polypeptide Chains. Let’s have a deep dive into the E. MS2 Genome, and hopefully try to decrypt it.
Reverse Engineering a Genome is a bit similar to reversing a real world piece of code, here is the methodology I’ll be following in order to understand the Genes within the E. MS2 Genome:
- First, we need to have access to the AA/NA Sequences: For this, I’ll be using the NCBI Database to get the DNA for the E. MS2 Virus.
- Extract Any Known Genes: A DNA is made of sequences of Genes, we will try and correlate any sequence in the DNA against the proteins database.
- Understand The Gene’s Functionality: Just like breaking down a big program into couple of functions/methods/classes. Understanding each object of these will help us conclude the general functionality of the program.
E. MS2 DNA Sequence
By going to the main NCBI page, and do a quick search using the name “Escherichia MS2”, this will land us on the following page which contains the full genome of the Virus.
This long pattern is made of a Nucleotide Acids sequence, it’s the alphabet of the DNA, it’s made of 4 characters:
- A for Adenine
- C for Cytosine
- G for Guanine
- T for Thymine
It’s so fascinating that these sequences hold every bit of information about you, me and every living thing in existence.
Our digital systems -including memory devices- work in a binary state, an existing voltage or zero voltage, an existing ampere or zero ampere, a High Logical Level or Low Logical Level, and we are able to make sense of this by grouping 8 bits together to form a byte.
Now imagine using the DNA as the storage medium (well, it is indeed *the* storage medium in our bodies and every living being), we have now 4-ary digital system, a quaternary-state system that’s made of 4 values, A, C, G or T. Each three of these states are grouped together to form a Codon, you can think of that as a Genetic Byte. Each gByte (Genetic Byte) is synthesized into an Amino Acid (not exactly, there is a step I jumped in here, the translation from DNA to mRNA, which is irrelevant to our context in this article), a chain of Amino Acids form a Peptide, a chain of Peptides form a Polypeptide, sometimes some contexts use Polypeptides interchangeably with Proteins.
In other words, a Genetic Byte is 6 bits, which makes the E. MS2 Genome 1143 Genetic Bytes, or 6858 digital bits, or almost 860 digital bytes. Awesome, isn’t it?
Some quick fun facts:
- Humans Genome is around 800MB, we only know way less than 100 MB of it.
- The Paris Japonica plant is one of the hugest Genome discovered, with around 40GB DNA size.
Okay back to our lovely E. MS2, Now we have access to the Genome of the Bacteriophage. To be more precise, to the DNA sequence of Nucleotide Acids base pairs.
Extracting Known Genes
Genes are made of Proteins/Polypeptides, which are chains of peptides made of chains of AAs, which are in turn synthesized from Codons sequences made of Nucleotides.
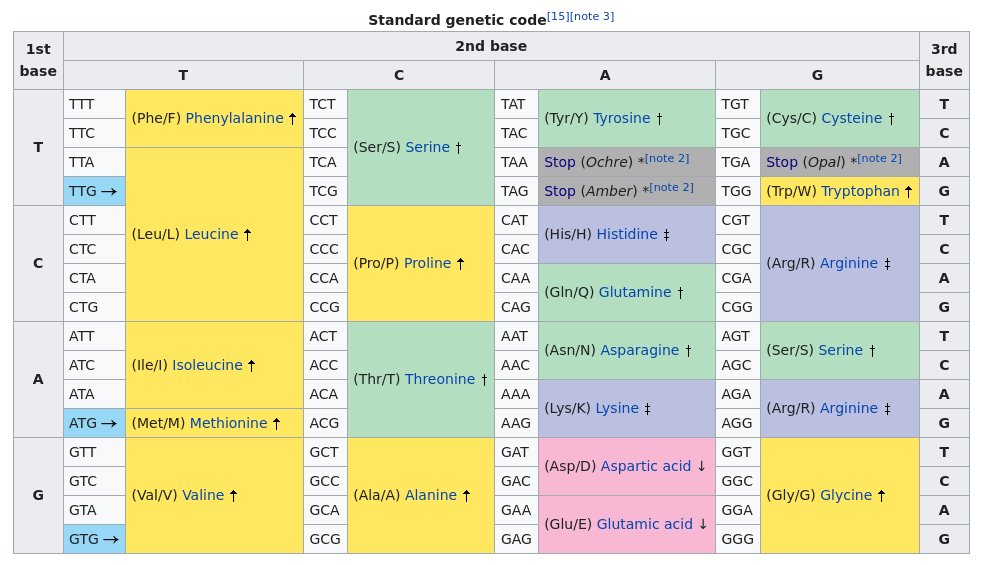
Let’s first try to translate the sequence of NAs we have to a sequence of CDS (Codons Sequence). This is done using the Codon Table.
Perfect! Now we are able to group every 3 Nucleotides together to form a single Codon, a piece of cake task for a Python script, no?
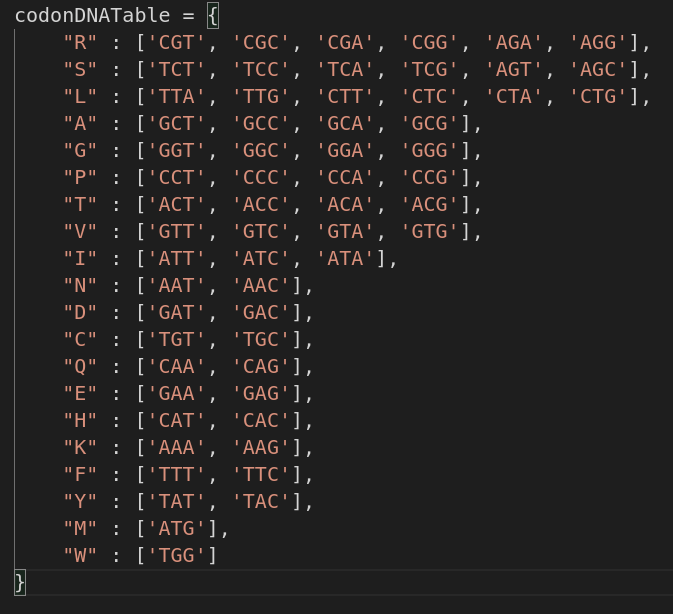
After the translation of each Codon in the NA sequence into its respective AA, we still can’t translate them right away into Polypeptides. Proteins vary in their lengths, and we are trying to reverse engineer an unknown sequence of AAs, how can we know that a given X number of AAs translate to Y Protein? One possible solution approach is Brute Force.
The idea is to keep reading the AAs until finding a complete Protein that translates into that sequence. Luckily, the NCBI BlastX Application does the heavy lifting for us, it takes a sequence of Nucleotide Base Pairs, translates them into the corresponding AAs, then tries to brute force its way through the resulting sequences to find any Protein that matches the given AA sequence in the provided Proteins Database.
You can use my script to facilitate querying the NCBI Blast App, it also stores the results in a text file for further investigation using text and regex processing utilities.
The results of the BlastX query is a long list of matches, statistical values, NAs and AAs. We are mainly interested in the Proteins that have a high match score. That is why I’m going to use the Linux utility “grep” to filter out the results and get only the proteins that fall in the 80th percentile match.
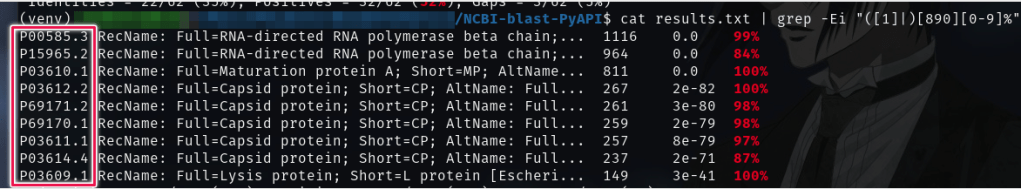
So far so good, we’ve got 4 types of Proteins at the top of the matches:
- Capsid Protein
- Lysis Protein
- Maturation Protein A
- RNA-directed RNA Polymerase Beta Chain
Understand The Gene’s Functionality
The highlighted area in the last screenshot shows the Protein Codes that will help us understand more about the functionality of the proteins, we only need to copy this code into the UnitProt Database, and find what the Biologists know so far about each protein.
- Capsid Protein P03612: The coat in which the Genome of the E. MS2 is stored and enclosed, it’s made of 178 capsids with triangular symmetry of 3 per protein. It also seems that it acts as a repressor when the polymerase finishes the synthesize of a whole replication of an E. MS2 cell. Fascinating!
- Lysis Protein P03609: Responsible for the lysis of the host’s cell membrane. Sounds like an “exfiltration” protein.
- Maturation Protein A P03610: Responsible for attaching the Capsid structure to the Bacteria’s Pilis (short hair-like bodies that are found at the surface of some Bacterias to help multidirectional swimming and maneuvring, as well as attach to their host). So that’s the protein responsible for infection.
- RNA-directed RNA Polymerase Beta Chain Protein P00585: A catalytic material for the replication of the Virus DNA inside the host.
__________________________________________________________________________
Results Evaluation and Conclusion
In order to evaluate how accurate our reversing journey was, we will need some references to compare our results against. Let us have a look at that Wikipedia page that contains links to the respective proteins in the NCBI database as well.
As far as we can tell, our results are decently accurate that we almost covered most of the information mentioned in the Wiki page Genome section, along with the names and functionalities of the genes and their synthesized Proteins. That gives us a rough idea on the efficiency of our methodology.